Resistance to Ciborinia camelliae within interspecific hybrids of Camellia
Denton-Giles M.1, Charvet D.2, Gordon T.R.3, Dijkwel P.P.1
1 Institute of Fundamental Sciences, Massey University, Palmerston North, 4442, NZ
2 Heartwood Nursery, Fort Bragg, CA 95437, USA
3 Department of Plant Pathology, University of California, Davis, CA 95616, USA
E-mail: P.Dijkwel@massey.ac.nz
Introduction
Ornamental Camellia shrubs are grown worldwide and are particularly renowned for their large, colourful winter blooms. However, the blooms of many ornamental Camellia hybrids are susceptible to infection by the fungal pathogen Ciborinia camelliae (Kohn). This pathogen exclusively infects the floral tissue of plants of the Camellia genus and is the most significant pest of ornamental Camellia plants (Kohn & Nagasawa, 1984). A successful infection leads to the rapid development of brown petal lesions and premature flower fall. Commonly referred to as camellia flower blight, this disease has been reported throughout the world, including the temperate regions of Japan, North America, New Zealand and Europe (Hara, 1919; Hansen & Thomas 1940; Taylor & Long, 2000).
Since the detection of Ciborinia camelliae in the USA in 1939 (Hansen & Thomas, 1940) concerted attempts have been made to control this pathogen. Previous methods have included the collection and destruction of infected blooms (Anzalone, 1959), the application of soil drenches, contact fungicides and systemic fungicides (Taylor & Long, 2000), as well as the inoculation of biological control agents (van Toor et al., 2005). Although several of these methods have shown promise in reducing the incidence of camellia flower blight, none have been able to stop the annual re-incursion of this disease into treated areas.
The recent discovery of natural resistance to C. camelliae in several species of the Camellia genus has revealed an opportunity to control this pathogen using conventional resistance breeding techniques (Taylor, 2004; Denton-Giles et al., 2013). Resistance appears to be prominent in Camellia species of section Theopsis, including C. lutchuensis, C. transnokoensis and C. fraterna. In the present work we aimed to determine whether resistance to Ciborinia camelliae is conserved within interspecific Camellia hybrids of section Theopsis origin.
Materials and methods
A total of 19 interspecific Camellia hybrids and 1 species were collected from Daniel Charvet’s population of Camellia hybrids (Table 1). Isolates of Ciborinia camelliae were collected from Heartwood Nursery, Fort Bragg, CA (coordinates 39° 25' 48" N and 123° 47' 8” W). Freshly collected fungal apothecia were inverted in 50 ml falcon tubes, and kept at room temperature to release their ascospores over a two day period. Ascospores were suspended in sterile H2O, quantified using a haemocytometer and diluted to a concentration of 8x105 spores/ml prior to petal inoculation.
The ascospore solution was sprayed onto the adaxial surface of detached Camellia petals using a 15 ml atomizer, with an estimated coverage of 500 to 1000 ascospores/cm2. Sterile H2O was used for all mock inoculations. Trays of inoculated and mock-inoculated petals were placed into plastic boxes and incubated in a growth cabinet at 21°C under a 12h/12h light/dark photoperiod at a photosyntheic photon flux density of 90 µmol m-2 s-1. Petals were removed for analysis at pre-determined time-points.
Images used for lesion analysis were taken using a Fujifilm FinePix Z digital camera. A total of 3 biological replicates were used to profile lesion development unless otherwise stated. ImageJ software (NIH, USA, http://rsb.info.nih.gov/ij/) was used to quantify lesion area at 16, 24, 48 hours post inoculation (hpi) using the ‘Colour Threshold’ tool. In addition, petals were assessed at 96 hpi for the presence of brown lesions.
An Olympus CH30 light microscope equipped with a Dinolite digital camera was used for microscopic analysis and image capture. For dual 3,3’-diaminobenzidine (DAB) (Sigma-Aldrich) and Trypan blue (Sigma-Aldrich) staining, 10 mm diameter discs of petal tissue were excised from live tissue two hours before their pre-determined assessment time-point, vacuum infiltrated with a solution of 1 mg/ml DAB pH 5.0 and dark incubated. At the time of assessment, discs were fixed and cleared in 3:1 ethanol: acetic acid and stained with 0.1% Trypan blue. For Trypan blue staining, discs were submerged in a solution of 0.1% (w/v) Trypan blue in lactophenol for 2 to 3 hours, washed twice in MilliQ H2O and stored in 30% (v/v) glycerol until required. Microscopic analysis was performed for each genotype by assessing 50 germinated ascospores at 24 hpi. Ascospores were considered to have germinated if their oval shape had become polarized or if primary hyphae were visible. Papilla formation was quantified by counting the number of interactions that produced a DAB-stained halo at the point of penetration. Lengthened primary hyphae were quantified by counting the number of interactions that produced a primary hypha with a length greater than its associated ascospore. Subcuticular hyphae were quantified by counting the number of interactions that gave rise to hyphae below the petal cuticle and, in turn, grew as far as the epidermal intercellular space. Subepidermal hyphae were quantified by counting the number of interactions that successfully gave rise to hyphae that were visible at a subepidermal focal plane.
The heat-map was generated from mean percentage values for each microscopic parameter using MeV software (Institute for Genomic Research, Rockville, MD; http://www.tm4.org/mev/). Papilla formation data were converted to papilla absence (inverse measurement of papilla formation) in order to conform to the high-to-low trend of the other heat-map data. The combined resistance score was calculated by ranking each Camellia genotype from 1 to 21 for each resistance parameter (i.e. A to E, Figure 3) and then calculating the sum of the combined ranking values. Mock inoculated controls were assessed for lesion development at 48 hpi. The software package R (R Foundation for Statistical Computing, Vienna, Austria; http://www.r-project.org/) was used for statistical analyses of the data. To compare significance between quantified microscopic resistance parameters, raw data from the Camellia ‘Nicky Crisp’ control was compared with other Camellia genotypes using multiple two-tailed Student’s t-tests. P values were adjusted for multiple testing using a false discovery rate correction (Benjamini et al., 1995).
Table 1. Descriptions of the interspecific hybrids and species of Camellia used in this study.
|
No. |
Parentage |
Cultivar names |
Mean bloom diameter (mm) |
|
1. |
(C. reticulata x C. japonica) x [C. reticulata x (C. saluenensis x C. transnokoensis) |
Heartwood ‘0216’ |
70 |
|
2. |
C. pitardii var. pitardii x C. japonica (Control) |
Camellia ‘Nicky Crisp’ |
63 |
|
3. |
C. pitardii var. yunnanica x (C. reticulata x [C. reticulata x C. fraterna])A |
Heartwood ‘9619’ |
63 |
|
4. |
(C. pitardii var. yunnanica x C. reticulata) x (C. japonica x C. transnokoensis) |
Heartwood ‘1113’ |
42 |
|
5. |
(C. reticulata x C. japonica) x (C. reticulata x [C. reticulata x C. fraterna]) |
Heartwood ‘0333’ |
65 |
|
6. |
(C. pitardii var. yunnanica x C. reticulata) x (C. saluenensis x C. transnokoensis) |
Heartwood ‘201’ |
90C |
|
7. |
(C. pitardii var. yunnanica x C. reticulata) x [(C. pitardii var. yunnanica x C. reticulata) x (C. japonica x C. fraterna)] |
Camellia ‘Avalon Sunrise’ |
68 |
|
8. |
C. pitardii var. yunnanica x (C. reticulata x [C. reticulata x C. fraterna])A |
Heartwood ‘9612’ |
72 |
|
9. |
C. pitardii var. yunnanica x (C. pitardii var. yunnanica x C. fraterna) |
Camellia ‘Nabb’ |
47 |
|
10. |
(C. pitardii var. yunnanica x C. yunnanensis) x (C. japonica x C. lutchuensis) |
Heartwood ‘1155’ |
68 |
|
11. |
(C. pitardii var. yunnanica x C. yunnanensis) x [(C. pitardii var. yunnanica x C. yunnanensis) x (C. japonica x C. lutchuensis)] |
Heartwood ‘1135’ |
73 |
|
12. |
[(C. pitardii var. yunnanica x C. reticulata) x (C. japonica x C. fraterna)] x (C. pitardii var. yunnanica x C. japonica) |
Heartwood ‘0376’ |
57 |
|
13. |
(C. pitardii var. yunnanica x C. reticulata) x (C. japonica x C. fraterna) |
Heartwood ‘77D’ |
45 |
|
14. |
(C. pitardii var. yunnanica x C. reticulata) x C. grijsii |
Heartwood ‘9860’ |
60C |
|
15. |
([C. pitardii var. yunnanica x C. reticulata] x C. forrestii) x C. japonica. |
Heartwood ‘0990’ |
82 |
|
16. |
(C. pitardii var. yunnanica x C. yunnanensis) x (C. japonica x C. lutchuensis) |
Heartwood ‘1139’ |
78 |
|
17. |
C. saluenensis x C. lutchuensis |
Camellia ‘Salute’ |
38 |
|
18. |
C. japonica x C. lutchuensis |
Camellia ‘Scentuous’ |
52 |
|
19. |
C. japonica x C. transnokoensis |
Camellia ‘Sweet Jane’ |
50 |
|
20. |
C. lutchuensis (Control)B |
Camellia lutchuensis |
27 |
|
21. |
C. lutchuensis (Control)B |
Camellia lutchuensis |
23 |
A Morphologically diverse offspring from the same original cross.
B C. lutchuensis individuals from two different localities.
C Data based on two biological replicates.
Results and discussion
Camellia ‘Nicky Crisp’ and C. lutchuensis petals were tested for their ability to promote and resist Ciborinia camelliae infection respectively. Disease-associated lesions were first visible on Camellia ‘Nicky Crisp’ tissue from 24 hours post inoculation (hpi) and had almost completely covered the petals by 48 hours (Figure 1A). In contrast, C. lutchuensis petals failed to develop disease-associated lesions, although petals had become partially dehydrated by 48 hours (Figure 1A’). Microscopic examination of inoculated C. ‘Nicky Crisp’ petal tissue showed fungal hyphae migrating from the ascospore to the intercellular space (Figure 1B). Hypha development on C. lutchuensis tissue was perturbed by the formation of plant papillae which are often associated with a localized burst of H2O2 (Figure 1B’). Papillae act to inhibit penetration of the plant cuticle by fungal hyphae and are characteristic of an induced-resistance response (Fan and Doerner, 2012).
Despite the major difference in locality, these results are consistent with what has been previously reported for Ciborinia camelliae-C. ‘Nicky Crisp’ and Ciborinia camelliae-C. lutchuensis interactions in New Zealand (Denton-Giles et al., 2013). The similarity in results between the two studies suggests that the fungal isolates used were not considerably different in their virulence. DNA sequencing showed that the fungal isolates from New Zealand and California used in these two studies were 99.8% identical in DNA composition across 2000 base pairs (data not shown).
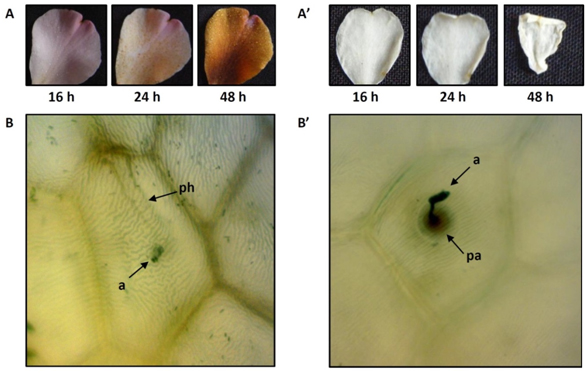 Figure 1. Visual examples of factors associated with disease development and plant resistance on susceptible Camellia ‘Nicky Crisp’ and resistant C. lutchuensis petals. Disease-associated lesions develop from 24 hours to 48 hours on C. ‘Nicky Crisp’ petals (A) but fail to develop on C. lutchuensis (A’). An ascospore germinates to produce an invasive subcuticular hyphae in C. ‘Nicky Crisp’ (B). Penetration of the petal cuticle is hampered by the formation of a papilla in C. lutchuensis petal tissue (B’). Symbols: a = ascospore; ph = primary hypha; pa = papilla. Magnification of image B and B’ is 400x.
Figure 1. Visual examples of factors associated with disease development and plant resistance on susceptible Camellia ‘Nicky Crisp’ and resistant C. lutchuensis petals. Disease-associated lesions develop from 24 hours to 48 hours on C. ‘Nicky Crisp’ petals (A) but fail to develop on C. lutchuensis (A’). An ascospore germinates to produce an invasive subcuticular hyphae in C. ‘Nicky Crisp’ (B). Penetration of the petal cuticle is hampered by the formation of a papilla in C. lutchuensis petal tissue (B’). Symbols: a = ascospore; ph = primary hypha; pa = papilla. Magnification of image B and B’ is 400x.
A total of 18 interspecific Camellia hybrids were chosen for resistance screening (Table 1). Each hybrid had one of the section Theopsis species C. lutchuensis, C. transnokoensis, C. fraterna, C. grijsii or C. forrestii as a contributing parent, although the total number of individual parents contributing to each hybrid varied from 2 to 6. To test the ability of these interspecific Camellia hybrids to resist Ciborinia camelliae, we quantified infection parameters associated with disease resistance (Figure 2).
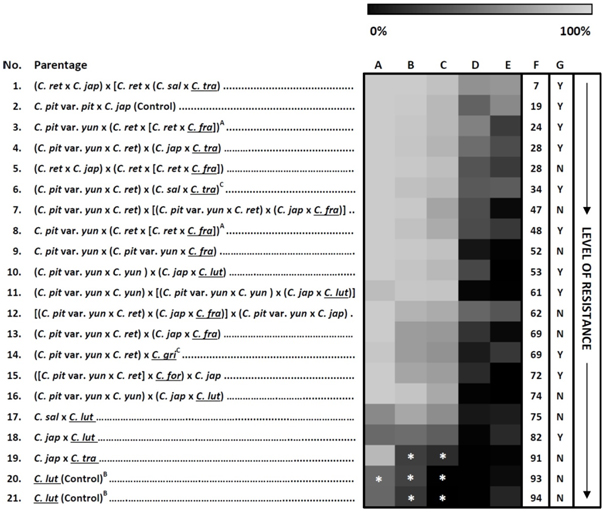

Figure 2. A graphical comparison of the Ciborinia camelliae resistance screening data across 19 interspecific hybrids and 1 species of Camellia (n=2). Column A= papillae absence, B = lengthened primary hyphae, C = subcuticular hyphal growth, D = subepidermal hyphae at 24 hours post inoculation (hpi), E = percent lesion area at 48 hpi, F = combined resistance score, and G = lesion development in mock-inoculated controls by 96 hpi (Y = yes, N = no). Tile colour coding represents mean percentage data of three biological replicates (unless specified), ranging from highly susceptible phenotypes (light gray) to comparatively resistant phenotypes (black). The displayed order of the Camellia genotypes is replicated in Table 1 and is based on the combined resistance score (F). Multiple two-tailed Student’s t-tests were applied to raw data for each independent column, comparing each genotype with the susceptible Camellia ‘Nicky Crisp’ (hybrid No. 2) genotype; * indicate values significantly less than Camellia ‘Nicky Crisp’ values at P = 0.1. The absence of asterisks represents no significant difference. Camellia species names are abbreviated as C. jap (C. japonica), C. pit var. pit, (C. pitardii var. pitardii), C. sal (C. saluenensis), C. pit var. yun (C. pitardii var. yunnanica), C. ret (C. reticulata), C. fra (C. fraterna), C. tra (C. transnokoensis), C. gri (C. grijsii), C. for (C. forrestii), C. lut (C. lutchuensis), C. yun (C. yunnanensis). All section Theopsis species are underlined. Superscript A: morphologically diverse offspring from the same original cross. Superscript B: C. lutchuensis individuals from two different localities. Superscript C: data based on two biological replicates.
Papillae were absent from the majority of the tested interactions (Figure 2, column A). Interestingly, the genotypes that scored low on the ‘papillae absence’ parameter all had C. lutchuensis within their parentage. Papillae production in C. lutchuensis has been previously reported and is thought to be a contributing factor to camellia flower blight resistance (Taylor 2004, Denton-Giles et al., 2013).
Resistance was also assessed in respect to the development of the fungus. Inhibition of fungal growth was measured by assessing the number of lengthened primary hyphae, the number of subcuticular hyphae and the number of subepidermal hyphae (Figure 2, columns B, C and D). Overall, fungal development was relatively unhindered in the tested interspecific hybrids, except for hybrid No. 19 (C. japonica x C. transnokoensis). The frequency of subepidermal hyphae was less than 2% for hybrid No. 19 (C. japonica x C. transnokoensis) and hybrid No. 16 (C. pitardii var. yunnanica x C. yunnanensis) x (C. japonica x C. lutchuensis) at 24 hpi, indicating that these two hybrids somehow slowed the development of the fungus during its establishment in the petal mesophyll tissue. The mechanism that inhibits hyphal growth in these genotypes is unknown. We speculate that plant antimicrobial secondary metabolites may contribute to the observed phenotype as the focal accumulation of antimicrobial compounds to infection sites has been previously reported as a hallmark of plant resistance (Underwood and Somerville, 2008).
Only 9 of the 21 genotypes had a mean lesion area average of less than 10% by 48 hpi (Figure 2, column E). C. lutchuensis petals also had a considerable degree (6 to 18%) of browning by this time, although it was difficult to determine whether this was due to the development of disease, due to a hypersensitive plant resistance response, or due to dehydration. Only hybrids No. 18 (C. japonica x C. lutchuensis) and No. 16 (C. pitardii var. yunnanica x C. yunnanensis) x (C. japonica x C. lutchuensis) failed to develop all enveloping disease-associated lesions by 96 hpi. Surprisingly, the mock-inoculated control of hybrid No. 18 (C. japonica x C. lutchuensis) did develop brown lesions, but it is unclear whether this was due to Ciborinia camelliae, or as a result of infection from an unidentified Camellia pathogen. Despite efforts to select uninfected blooms for resistance screening, it is plausible that many of the blooms were exposed to C. camelliae ascospores prior to sampling.
Hybrids were ranked based on their resistance score (Figure 2, column F). Interspecific hybrids with large resistance scores tended to have C. lutchuensis as their contributing Theopsis parent, with the exception of hybrid No. 19 (C. japonica x C. transnokoensis). Furthermore, the highest scoring C. lutchuensis hybrids had higher dosages of C. lutchuensis within their parentage and were often derived from only two parents. Hybrids that had C. lutchuensis within their parentage but also had several other contributing parents tended to be more susceptible, with the exception of hybrid No. 16 (C. pitardii var. yunnanica x C. yunnanensis) x (C. japonica x C. lutchuensis), which was the only interspecific Camellia hybrid that failed to develop disease. Images of the more resistant C. lutchuensis hybrids are shown in Figure 3, together with some of the more susceptible interspecific hybrids.
Interestingly, hybrids with identical or similar parentage did not respond in the same way to Ciborinia camelliae infection. The strong resistance phenotype demonstrated by hybrid No. 16 (C. pitardii var. yunnanica x C. yunnanensis) x (C. japonica x C. lutchuensis) was not conserved in hybrid No. 10 (C. pitardii var. yunnanica x C. yunnanensis) x (C. japonica x C. lutchuensis). Furthermore, hybrids that were derived from exactly the same parent plants (i.e. hybrid Nos. 3 and 8) also scored differently, although they were both ultimately susceptible to Ciborinia camelliae. These results suggest that although parentage is relatively conserved in these hybrids, each interspecific hybridization event is unique and ultimately determines the genetic outcome of the offspring. In addition to recombination, complex genetic rearrangement can occur during interspecific hybridization that can lead to changes in gene expression (Johnson, 2008). These changes could conceivably influence disease resistance as this trait is under genetic control.
The majority of the interspecific Camellia hybrids tested were susceptible to Ciborinia camelliae. However, observations of lesion development at 16, 24, 48 hpi indicated that the rate at which disease developed varied between hybrids. Furthermore, several hybrids that exhibited a low incidence of lesion development at 48 hpi went on to develop lesions by 96 hpi, including hybrid No. 15 ([C. pitardii var. yunnanica x C. reticulata] x C. forrestii) x C. japonica, hybrid No. 17 (C. saluenensis x C. lutchuensis) and hybrid No. 19 (C. japonica x C. transnokoensis). This ‘delayed susceptibility’ phenotype offers an alternative strategy for reducing camellia flower blight symptoms on plants. If this trait was to be selected for in hybrids that have short-lived blooms, disease could potentially be abscised from the plant before it had a chance to negatively affect the aesthetic value of the bloom. However, this strategy may not reduce the incidence of Ciborinia camelliae in the wild, as it can persist on abscised blooms (Kohn & Nagasawa, 1984).

Figure 3. Examples of blooms from interspecific Camellia hybrids that are highly susceptible to (A to C), and are resisting (D to F) Ciborinia camelliae infection. Image A = No. 1 (C. reticulata x C. japonica) x [C. reticulata x (C. saluenensis x C. transnokoensis), B = No. 3 C. pitardii var. yunnanica x (C. reticulata x [C. reticulata x C. fraterna]), C = No. 4 (C. pitardii var. yunnanica x C. reticulata) x (C. japonica x C. transnokoensis), D = No. 16 (C. pitardii var. yunnanica x C. yunnanensis) x (C. japonica x C. lutchuensis), E = No. 17 C. saluenensis x C. lutchuensis, F = No. 18 C. japonica x C. lutchuensis.
Conclusion
The data presented here demonstrate that Camellia lutchuensis derived hybrids were the most resistant to infection by Ciborinia camelliae, with the exception of one C. transnokoensis-based hybrid. However, for all but one interspecific Camellia hybrid (hybrid No. 16), the levels of natural resistance were not sufficient to prevent the development of camellia flower blight. These data support the use of C. lutchuensis in future breeding strategies that promote Ciborinia camelliae resistance in ornamental Camellia hybrids. We propose crossing current C .lutchuensis hybrids back to C. lutchuensis in order to increase the C. lutchuensis genetic dosage within interspecific Camellia hybrids.
Acknowledgements
This work was supported by the New Zealand Camellia Memorial Trust scholarship and Palmerston North Jaycee Trust Travelling Scholarship. We thank Sharon Kirkpatrick of UC, Davis’ plant pathology department for technical assistance.
References
Anzalone, L. J. 1959. Studies on flower blight of Camellia and Cercospora leaf spot of Photinia serrulata. Doctoral dissertation, Louisiana State University, Baton Rouge, USA.
Benjamini, Y., and Hochberg, Y. 1995. Controlling the false discovery rate – A practical and powerful approach to multiple testing. Journal of the Royal Statistical Society. Series B, Statistical methodology 57: 289-300.
Denton-Giles, M., Bradshaw, R. E., and Dijkwel, P. P. 2013. Ciborinia camelliae (Sclerotiniaceae) induces variable plant resistance responses in selected species of Camellia. Phytopathology 103: 725-732.
Fan, J., and P. Doerner. 2012. Genetic and molecular basis of nonhost disease resistance: complex, yes; silver bullet, no. Current Opinion in Plant Biology 15(4): 400-406.
Hara, K. 1919. On a Sclerotinia disease of camellia. Dainippon Saurinkwaiho 436: 29-31.
Hansen, H. H., and Thomas, H. E. 1940. Flower blight of Camellias. Phytopathology 30: 166-170.
Johnson, N. (2008) Hybrid incompatibility and speciation. Nature Education 1(1): 20.
Taylor, C. H., and Long, P. G. 2000. Review of literature on Camellia flower blight caused by Ciborinia camelliae. New Zealand Journal of Crop and Horticultural Science 28: 123-138.
Taylor, C. H. 2004. Studies of Camellia flower blight (Ciborinia camelliae Kohn). Doctoral dissert. Massey University, Palmerston North, New Zealand.
Van Toor, R. F., Jaspers, M. V., and Stewart, A. 2005. Effect of soil microorganisms on viability of sclerotia of Ciborinia camelliae, the causal agent of Camellia flower blight. New Zealand Journal of Crop and Horticultural Science 33: 149-160.
Underwood, W., and S. C. Somerville. 2008. Focal accumulation of defences at sites of fungal pathogen attack. Journal of experimental botany 59(13): 3501-3508.
Web design by Tribal Systems
