A flavonol synthase gene related to yellow flowers in Camellia nitidissima
Li Ji-yuan 1* Zhou Xing-wen 1,2 Fan Zhengqi 1 Yin Heng-fu 1* Sun Ying-kun 1 Li Xin-lei 1 Chen Yue 1
1.The Research Institute of Subtropical Forestry, Chinese Academy of Forestry, Fuyang, 311400, Zhejiang, China
2.Yulin normal university, Guangxi Zhuang Autonomous Region, Yulin 537000, Guangxi, China
1 Introduction
Flower color is one of the most important characters for ornamental plants,especially in customer selection among products in the floriculture market(Karen et al.2002;To et al. 2006). It is known that coloration of flowers is usually controlled by three classes of pigments: flavonoids, arytenoids and betalains (Tanaka et al. 2008).Generally, flavonoids and carotenoids are the widest pigments in plants because betalains only exist in few species from several generals such as Amaranthus, Celosia, Gomplrena, Iresine, etc(To et al. 2006; Tanaka et al.2008).In plants, flavonoids are the most common pigments which contribute to a range of flower color from yellow to purple(To et al.2006)in addition to they play a crucial role in resistance(Treutter2005), anxin transport(Peer et al. 2007) and signaling(Winkel-Shirley,2001). Flavonoids which include anthocyanins, flavones, isoflavones, flavonols etc (Taylor &Grotewold.2005;Grotewold.2006;Lepiniecet al. 2006; Subramanian et al. 2007) are extensive studied. The pathway of flavonoid biosynthesis has been well understood and the genes encoding the biosynthetic enzymes have been identified (Nieslenet al.2002; Tanaka et al.2008; Buer et al.2010).
Camellia nitidissima is a famous ornamental plant which is known as ‘the Queen of camellias’ due to its golden flowers (Chang 1979;1998; Gao et al.2005). It is an important parent species for breeding the yellow camellias due to that the color of camellia’s flowers is usually red, purple, pink and white, and the yellow is quite rare. However, the camellias with good yellow flowers have not appeared through many years’ efforts of traditional breeding(Chen 1987; Parks C R 2000; Nishimoto S et al. 2004). The appearance of transgenic plants with modified colors such as gerbera(Elomaa et al.1993), Petunia(van der Krol et al. 1988), lisianthus (Deroles et al.1998;Nielsen, 2002),violet carnation(Fukui et al. 2003), Torenia(Fukusaki et al.2004) and blue rose(Katsumoto et al.2007) shows that it is possible to get the new flower color which can’t obtained by traditional breeding through genetically modification.
The previous researches indicate that the main pigments of C. nitidissimaare quercetin-7-O-β-D-glucoside(Qu7G)and quercetin-3-O-β-D-glucoside(Qu3G) (Miyajima et al.1985; Peng et al. 2011). Quercetin(Qu)belongs to flavonols, which are among the most abundant flavonoids in plants and are usually found in the form of mono-, di- or triglycosides (Winkel-Shirley,2001;Bure etal.2010;Stracke et al. 2010).The biosynthesis of flavonols is one of the branches of flavonoids pathway and the flavonols are originated from the dihydroflavonols which are catalyzed by the enzyme flavonol synthesis (FLS) (Winkel-Shirley,2001;Nieslen et al.2002).Flavonols are usually act as copigment and can affect the flower color(Yoshitamaer al. 1992;Aida et al. 2000), and are responsible for the yellow petal color of Lathyrus chrysanthus(Markham et al.1994). The FLS gene has been cloned and used to alter the flower color successfully such as petunia(Holton et al.1993; Tsuda et al. 2004),lisianthus(Nielsen et al. 2002).
We have cloned an FLS gene (CnFLS, Genbank accession number:JF343560.1) from the petals of C.nitidissima, but the relations between this gene and the flower color is unknown. In order to clarify the biological role of CnFLS in vivio, expression vectors with sense CnFLS, RNA interference (RNAi) segments of CnFLS were constructed respectively in this study. The transformation of model plants Nicotianatabacum byAgrobacterium-mediated gene transfer was also performed. The results show that the transgenic plants with CnFLS gene grew normally and the flower color changed greatly compared to the wild type plants. The flower color of transgenic plants was inherited stably over two generations.
2 Materials and methods
2.1 Plant Material
The leaves of N. tabacum were chosen from the 1-month-old plants which were cultivated in the sterile culture bottle containing 35 mL of solid Murashige and Skoog (MS) medium.
2.2 The obtaining of T0 and T1 plants of transgenic N. tabacum
Leaves were cut to about 1cm2 and co-cultivated with Agrobacterium tumefaciens EHA105 cells carrying the pCAMBIA1300CaMV35S-CnFLSvector and pCAMBIA1300CaMV35S-CnFLS RNAi vector respectively according to the methods of An et al(1986).The binary vector pCAMBIA1300 carries the hygromycin B resistance gene under the CaMV35S promoter for selection of transgenic plants. After co-cultivation for 3days in dark room, EHA105 cells were washed off by sterile water and the leaves were grown on a callus induction MS medium containing 0.5mg L-1 6-benzyladenine, 20mg L-1hygromycin B and 200mg L-1Timentin. The callus induction medium was replaced every 2 weeks until the shoots emerged. When the shoots were 2cm high they were transferred to another MS medium containing0.1 mg L-1 α-naphthaleneacetic acid, 20mg L-1hygromycin B and 200mgL-1Timentin until roots grew. The rooted plants were then cultivated in a greenhouse at 25°C.After the insertion of foreign gene in transgenic individuals was analyzed by polymerase chain reaction (PCR), the plants were cultivated to flower. The T1 seeds from the plants whose flower color differenced to the control were obtained after the cross-pollination prevented (Qiu et al. 2011) and then were selected by germinating on the MS medium containing 20 mg L-1hygromycin B. The T1 plants were grown to maturity and used for further experiments.
2.3 DNA Extraction and PCR Analysis
Genomic DNA was isolated from young leaf tissues using Column Plant DNA out Kits (TIANDZ, China). The forward primer 5'-GCGTCTGCTGCTCCATACA-3' and the reverse primer 5'-TGACATTGGGGAGTTTAGCG-3' which are specific for 0.5kb fragment of hygromycin gene were used to detect the recombinant DNA. The PCR system involved 150ng of total DNA, 1×PCR buffer for KOD FX, 0.4mM dNTP, 0.3μM primers, 1U of KOD FX(TOYOBO, Shanghai, China) and the PCR conditions as followed: pre-denaturation at 94°C for 2min, denaturation at 98°C for 10sec, annealing at 55°C for 30sec, extension at 68°C for 30sec, 30cycles later a 7min extension followed. After the amplification, the PCR product was detected by the electrophoresis on a 1.0% agarose gel. The positive transgenic plants were chosen for further tests.
2.4 Southern blotting
About 20μg genomic DNA was digested with restriction endonuclease HindIII (MBI Fermentas, Canada) at 37°C for 16 hours, electro phoretically separated on a 1.2% agarose gel(Sambrooket al. 1989) and transferred to a positively charged nylon membrane().The DNA was fixed on the membrane by baking at 120°C for 30min. The preparation of CnFLSprobe, per hybridization, hybridization and immunological detection were all performed according to the protocol of DIG-High Prime DNA Labeling and Detection starter KitⅠ(Roche, USA).
2.5 Real-time fluorescence quantitative PCR
Total RNA was extracted from the leaves of transgenic plants and control plants using the Column Plant RNAout2.0 kit and treated with Column DNA Erasol (Beijing Tiandz gene Technology Company, Beijing, People’s Republic of China) to avoid the DNA contamination(Zhu et al. 2011). The purified total RNA was reverse transcribed using oligo (dT) primer and PrimeScript® RT reagent Kit(TAKARA, Japan) according to the manufacturer’s instructions. The gene-specific primers of PCR amplification for CnFLSand Actinc DNAs were designed. To investigate the expression of CnFLS, the real-time PCR was performed with ABI PRISM 7300 Real-Time PCR System (USA) and SYBR Premix Ex Taq (TAKARA, Japan). Amplification occurred in a two-step procedure: denaturation at 95°C for 30 s and followed 40 cycles with denaturation at 95°C for 5s, 60°C for 31s. After completion of the amplification steps, the melting curve was determined for each analysis and the data were analyzed with the 2-ΔΔCT method (Livak, et al. 2011).
2.6 Pigment analysis
The pigments of flavonoids were extracted from the fresh colored coronal (about 50mg) of N. tabacum flowers which were blooming (Sun et al.2011) by the 2mL of MeOH:H2O: AcOH: TFA (70:27:2:1, V/V) at 4°C for 24 hours avoiding light. The Extracted solvent was filtrated by 0.22μm microporous membrane before High-performance liquid chromatography (HPLC) analysis. HPLC analyses were carried out on an Inertsil ODS-SP (4.6×250mm,5μm) column in LC-20AT(Shimadzu, Japan) and the temperature of column was 30°C. The solvent A of 1.5% H3PO4 and B of HCOOH: MeCN: THF: H3PO4: H2O(25:20:5:1.5:48.5) were applied as a gradient, changing line from 82%A and 25%B(0min) to 18%A and75%B(40min) and the flow rate was 0.8mL/min. 20μL of extracted solvent was injected and analyzed in one sample. The chromatogram was recorded at the absorption spectra 525nm for anthocyanthin (cyanidin-3-O-rutinoside,Cy3R)(Sun et al. 2011) and 372nm for flavonols Qu3G, Qu7G and quercetin(Parks et al. 1988). The peaks representing the Cy3R and the flavonols were confirmed via their own characteristic absorption spectra. The integrals were used for quantification of pigments by comparison with standard curves for Cy3R, Qu3G, Qu7G and quercetin measured at the same wave lengths and conditions. The standard samples of Qu3G, Qu7G, and quercetin for HPLC were from Sigma (Germany) while Cy3R was from Wako (Japan). All the experiments and the measurements were performed at least three times independently. The data were analyzed by the software of SPSS17.0 version.
3 Results
3.1 Transformation and generation
The expression binary vectors of CnFLS gene and RNA interferrence of CnFLS gene used in this study were shown as figure.1-A, B. Mediated by Agrobacterium EHA105, the gene CnFLS and RNAi of CnFLS were transformed into tobacco plants respectively (Fig.2).After co-cultivation for 15-25days, the resistant tiny shoots appeared. The green shoots about 2cm high were transferred to the rooting medium containing hygromycin and the root system formed about 1 month later. Putative transgenic lines of fifty sense CnFLS and forty lines ofCnFLSRNAi were obtainedf rom hygrocymin resistant shoots. When flowered, the color of 10 lines with sense CnFLS gene and 6 lines of CnFLSRNAi gene changed compared with wild type plants. The T0 seeds of the transgene plants whose color changed were collected. The survival of T1 seeds showed an approximately ration of 3:1 when selected by hygromycin and grew to flower.
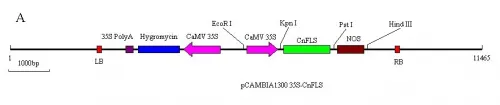
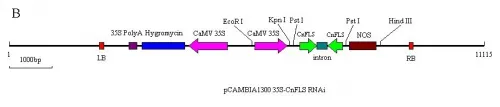
Fig.1A) The schematic expression structure of the T-DNA region of pCAMBIA1300 35S-CnFLSvector.
B) The schematic expression structure of the T-DNA region ofpCAMBIA1300 35S-CnFLSRNAivector. LB: left border of pCAMBIA1300, 35S Poly A: poly A of CaMV 35S, Hygromycin: Hygromycin B phospho transferose gene CaMV 35S: promoter of CaMV 35S, CnFLS: FLS gene from C.nitidissima, intron: the 1st intron of GA20 oxidase from potato, NOS: nopaline synthase terminator, RB: right border of pCAMBIA1300
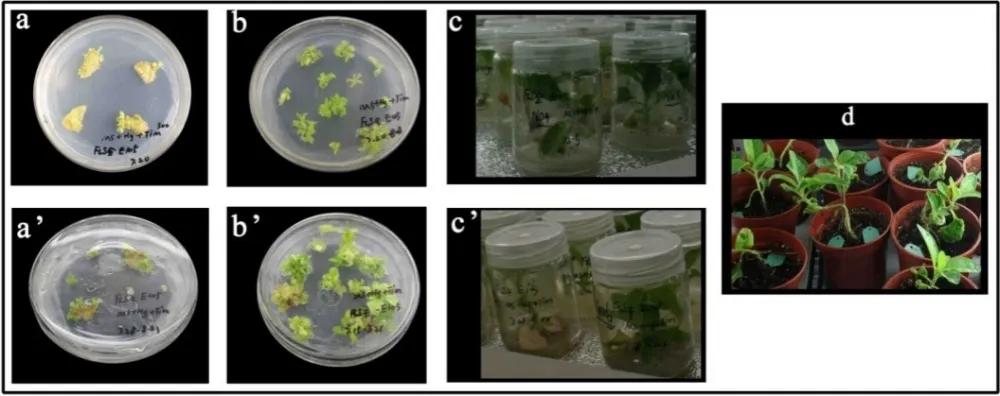

Fig.2 Transformation of N.tabacum and the plant regeneration. a: induced shoots of sense CnFLS, b:transplanted shoots of sense CnFLS, c: rooting plants of sense CnFLS, d:transplant of transformants, a’: induced shoots of RNAiCnFLS, b’:transplanted shoots of RNAiCnFLS, c’: rooting plants of RNAiCnFLS.
3.2 Identification of transformants
Transgenic lines TF1, TF4, TF31of sense CnFLSgene(Fig.3-A) and transgenic lines Tf1, Tf23, Tf35of RNAiCnFLS gene(Fig.3-B) had significantly different flower color from the untransformed control plants. The plants of TF1, TF4 and TF31 showed white to cream color while the plants of Tf1, Tf23 and Tf35 showed reddening effects in the flowers. The electrophoresis of agarose gel showed that a 500bp fragment of hygromycin gene was amplified from both the plasmids and the transgenic lines while no corresponding fragment from non-transgenic plants (Fig.3-C). The results of Southern blotting indicated that a single copy of sense CnFLS gene was integrated into the genomic DNA of these positive transgenic lines and the same results to the positive lines of RNAiCnFLStransformants(Fig.3-D).
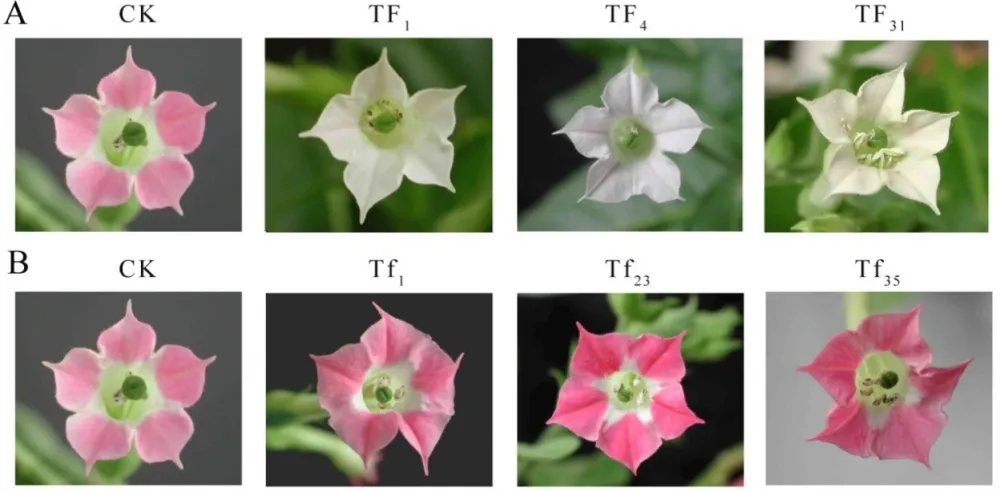

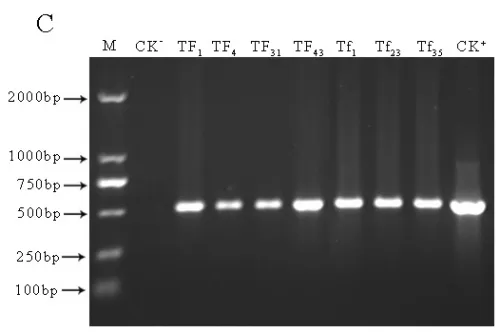
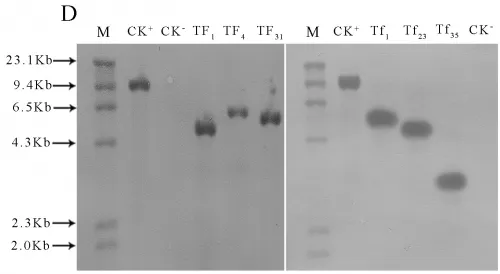
Fig.3(A)Comparison of the flower color of untransformed tobaccos and flowers from transgenic tobaccos of sense CnFLS gene.CK: flower of untransformed tobacco, TF1, TF4 and TF31: transgenic lines of sense CnFLS gene.(B)Comparison of the flower color of untransformed tobaccos and flowers from transgenic tobaccos of RNAiCnFLS gene.CK: flower of untransformed tobacco, Tf1, Tf23 and Tf35: transgenic lines of RNAiCnFLS gene.(C)Detection of hygromycin by PCR. M: molecular weight marker DL2000,CK-: negative control (wild type tobaccos),lanes TF1, TF4,TF31, TF43,Tf1, Tf23, and Tf35: putative transgenic lines, CK+: positive control (plasmid of pCAMBIA1300).(D) Southern blot analysis of transformed plants.M:λ-Hind III digest DNA Marker, CK+: positive control (plasmid of pCAMBIA1300), CK-: negative control (wild type tobaccos),lanes TF, TF4,TF31: transgenic lines of sense CnFLS,Tf1, Tf23 and Tf35:transgenic lines of RNAiCnFLS gene.
3.3 The expression analysis of CnFLS gene
Real-time quantitative PCR were performed with the total RNAs extracted from the leaves of transgenic tobacco linesTF1, TF4, TF31, Tf1, Tf23, Tf35 and untransformed plants which cultivated in greenhouse under the same normal condition. The results of expression level of FLS gene showed significant difference between transgenic plants and untransformed (Fig.4). In the transgenic lines of sense CnFLS, the expression level was at least 7 times higher than the wild type and the TF1line had the highest expression level which was almost 14 times of that of untransformed tobaccos. The results indicated that the sense CnFLS gene over expressed successfully under the control of CaMV 35S promoter in transgenic plants. The expression levels in RNAiCnFLS gene transformed lines were all lower than that of non-transgenic plants. The line of Tf23 had the lowest expression level which was about 0.2 times of that of wild type tobaccos. These data showed that the expression of FLS gene in transgenic tobaccos was inhibited to a great degree by the expression of RNAiCnFLS gene.
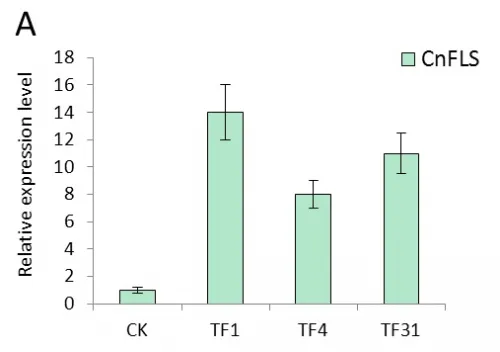
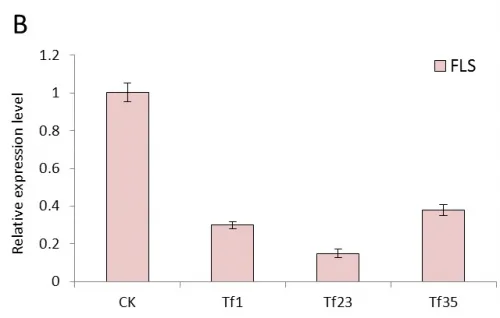
Fig.4 The expression analysis of CnFLS gene in the transgenic plants by Real-time PCR.(A): The expression level of transgenic plants of sense CnFLS gene. CK: negative control plants (wild type), TF1, TF4,TF31: transgenic lines of sense CnFLSgene. (B): The expression level of transgenic plants of RNAiCnFLS gene. CK: negative control plants (wild type),Tf1, Tf23 and Tf35: transgenic lines of RNAiCnFLS gene.
3.4 Pigments analysis by HPLC
The main pigment component in flowers of tobacco is cyanin-3-O-rutinoside(Cy3R) which is more than 90% of all anthocyanins (Aharoni et al. 2001). It is known that cyanin and Qu share the compound dihydroquercetin as the common precursor which can be catalyzed by FLS and dihydroflavonol-4-reductase (DFR) in the biosynthesis pathway(Tsuda et al. 2004, Grotewold. 2006, Stracke et al. 2007, 2010).The content of Cy3R, Qu3G, Qu7G and Quin the flower of transgenic and untransformed tobaccos were analyzed by HPLC(Fig.5-A, B, C, D).
The content of Cy3R in the transgenic lines of TF1, TF4 and TF31 decreased significantly compared with the control plants(wild type) (Fig.5A).The content of Cy3R in TF1 was about 0.05 times of that in wild type tobaccos and was lower than that of TF4 and TF31. As to the transgenic lines of Tf1, Tf23 and Tf35, the content of Cy3R was far higher than the non-transgenic plants (Fig.5B). The content of Cy3R in Tf23 was nearly 2 times higher than that of wild types.
Analysis of HPLC showed that the content of Qu increased in trans-sense CnFLS gene lines TF1, TF4, TF31 and decreased in the trans-RNAiCnFLS gene lines Tf1, Tf23 and Tf35. The highest content of Qu in TF1 was about 2 times of that in control tobaccos while the lowest was in Tf1(Fig.5C, D).
The contents of Qu7G and Qu3G which derived from the flavonol Qu were also analyzed by HPLC. The result indicated that the content of these two compositions increased in the transgenic lines of TF1, TF4, TF31 and decreased in Tf1, Tf23 and Tf35. Fig.5C showed that the content of Qu7G in TF1 was about 2 times of that in control plants. The most significant change was the content of Qu3G in TF1 which was nearly 3 times higher than the wild tobaccos. Fig.5D showed that the content of Qu7G and Qu3G decreased unobviously compared with the untransformed plants and Tf23 had the lowest contents of these two compositions among the three trans-RNAiCnFLS gens lines.
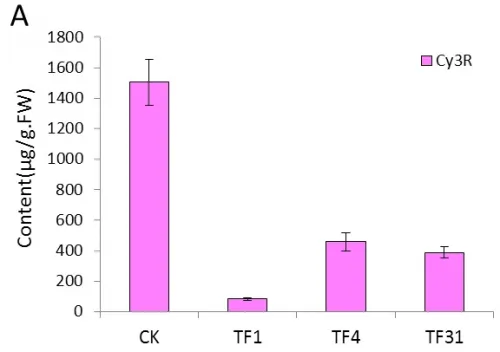
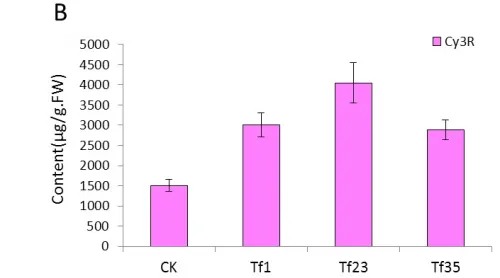
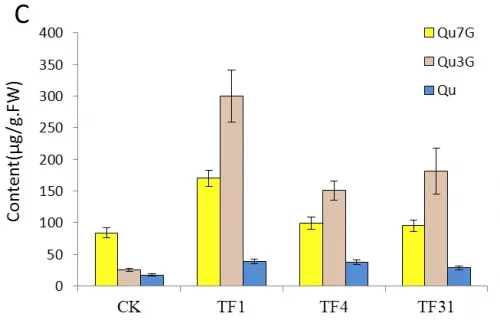
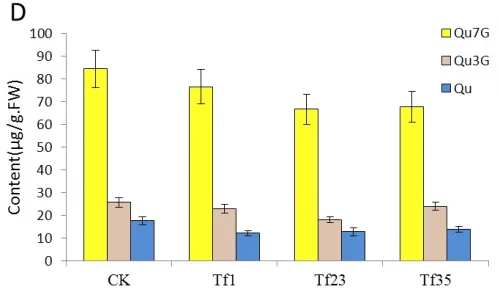
Fig.5 Comparsion of pigments contents in the flowers among the sense CnFLStransformed, RNAi CnFLS transformed and wild type tobaccos. (A) The Cy3R content analysis between the transgeinc lines of sense CnFLS gene and wild type tobaccos.CK: negative control plants (wild type), TF1, TF4, TF31: transgenic lines of sense CnFLS gene.(B)The Cy3R content analysis between the transgeinc lines of RNAi CnFLS gene and wild type tobaccos.CK: negative control plants (wild type), Tf1, Tf23 and Tf35: transgenic lines of RNAi CnFLS gene. (C)The Qu7G, Qu3G and Qu contents analysis between the transgeinc lines of sense CnFLS gene and wild type tobaccos.CK: negative control plants (wild type), TF1, TF4,TF31: transgenic lines of sense CnFLSgene.(D)The Qu7G, Qu3G and Qu contents analysis between the transgeinc lines of RNAi CnFLS gene and wild type tobaccos.CK: negative control plants (wild type), Tf1, Tf23 and Tf35:transgenic lines of RNAi CnFLS gene.
4 Discussion
Generally, the effects of one gene in transgenic plants can be studied from two aspects. The one is overexpression and the other is inhibition the expression of the gene by the method such as transferring the antisense gene, cosupression or RNAi etc.RNAi become more popular to the researcher because of its efficient, specific and steady characters(Nakamura et al.2006, Nakatsuka et al.2008,Kamiishi, et al.2011).In this study, sense CnFLS gene and RNAiCnFLS gene were successful transformed into tobaccos through the recombinant vectors pCAMBIA1300 35S-CnFLS and pCAMBIA1300 35S-CnFLSRNAi respectively.
Our results indicated clearly that the transgenic lines of sense CnFLS gene and RNAiCnFLSgene were obtained via the molecular identification and phenotype. Real-time PCR demonstrated that the CnFLS gene overexpressed in transgenic lines of sense CnFLS gene. The transgenic lines of sense CnFLS gene displayed white to faint yellow color which shows great difference to the pink of control plants. HPLC analysis showed that the content of Cy3R decreased significantly while the flavonols of Qu, Qu3G and Qu7G increased greatly. These results confirmed that the CnFLS gene expressed and played an important effect on the flower color. The enzyme FLS usually catalyzes the dihydrofavonol to flavonol which is a limb pathway in flower pigment biosynthesis(Nielsen et al, 2002; Davies et al. 2003,Grotewold 2006). It is the CnFLS utilize more dihydroquercetin as substrate to produce quercetin that the formation of Cy3R was limited, which may be the reason of the white flower color of the transgenic lines. However, the CnFLS can’t catalyze all the substrates that the Cy3R which is the production of DFR was detected by HPLC. That is why the transgenic lines of TF4 showed a little red color.
As to the transformants of RNAiCnFLS gene, the flower color indicates a trend from pink to deep red. The content of Cy3R which originated from dihydroquercetin increased 2 times that of control plants. The results are similar to the findings of Holton et al. (1993).They introduced an antisense FLS gene from Petunia hybrida intoNicotianata bacumand changed the flower color from pink to red. The quercetin and its glycosides were detected in this study indicates that the expression of FLS was not inhibited completely by the interference of CnFLS. Previous researches show that FLSis a member of multigene family in Arabidopsis(Pelletier et al. 1997) and lisianthus(Nielson et al. 2002), which may be true of tobaccos. That the expression of RNAiCnFLSspecifically suppressed the homologous gene but the expression of other members in this super family may result the biosynthesis of quercetin. The results that the loss expression of FLS led to an overall increase of anthocyanin are not in accordance with the findings of lisianthus which only accumulated the dihydroflavonols in the antisense FLS plants (Nielson et al. 2002). The main reason of the difference between the two species may be the substrate competition of FLS and DFR in tobaccos. This indicates that using RNAi, antisense or sense DFR may obtain the flowers with more anthocyanin.
The colorless flavonols are abundant in many flower petals and usually looked as a copigment in flower color because of its planar moleculars(Holton et al. 1993, Brouillard et al. 1993, Devies et al.2003, Nielson et al. 2002).However, the yellowish flower color of sense CnFLStransgenic tobaccos accumulated more flavonol glycosides rather than flavonol with the greatly decrease of Cy3R content. This might suggest that the flowers may show yellow color when the ratio of flavonol glycosides and anthocyanin is suitable and the content of flavonol glycosides is sufficient.
In conclusion, we obtained tobacco plants with novel flower color by modifying the flavonol synthesis. The expression of sense CnFLSresulted in the content increase of Qu and its glycosides Qu7G, Qu3G, the flower color showed fain yellow color at the same time. In contrast, the expression of RNAiCnFLSpromoted the accumulation of anthocyanin, Cy3R and the decline of Qu and its glycosides, the flower color changed into deep red. From these results we may conclude the CnFLS mainly catalyze the dihydroquercetin and produce Qu. This may be the reasonable explanation of that why plenty of Qu3G and Qu7G accumulate in the petals of C.nitidissima. This study indicates that the CnFLS play an important role in flower color determination and it can be used to bleach or deepen the flower color by transgenic technology to breed the new cultivar of ornamental plants.
Acknowledgments:
This work was supported by the National Key Twelfth-Five Science and Technology Program (2012BAD01B0703), International Cooperation Project of China (2011DFA30490), and Zhejiang Key Flower Breeding Program (2012C12909-6) as well as Basic research fund for Central public Scientific Research Institute (RISF6141).
Literature cited
[1] Aharoni A, De Vos CHR, Wein M, Sun ZK, Greco R, Kroon A, Mol JNM, O’Connell AP (2001). The strawberry FaMYB1 transcription factor suppresses anthocyanin and flavonol accumulation in transgenic tobacco. PlantJ28:319-332
[2] Aida R, Yoshida K, Kondo T, Kishimoto S, Shibata M(2000)Copigmentation gives bluer flowers on transgenic torenia plants with the antisense dihydroflavonol-4-reductase gene. Plant Science160:49-56
[3] An G, Watson BD, Chiang CC (1986) Transformation of tobacco, tomato, potato and Arabidopsis thaliana using a binary Ti vector system. Plant Physiol 81:301-305
[4] Buer C S, Imin N,Djordjevic M A (2010) Flavonoids: New Roles for Old Molecules.Journal of Integrative Plant Biology 52 (1):98-111
[5] Brouillard R, Dangles O. (1993) Flavonoids and flower colour. In: Harborne J.B. (ed.),The Flavonoids: Advances in Research Since 1986. Chapman & Hall, London, pp. 565-587.
[6] Chang H D(1979)Chrysantha, a section of golden camellias from cathaysianflora.Journal of Zhongshan University 3:69-74
[7] Chang H D, Ren S X (1998)Theaceae.In:Flora ReipublicaePopularisSinicae.Beijing:Science Press.49(3):87-133
[8] Chen J Y (1987) Fourteen years breeding of golden camellia. Journal of Beijing Forestry University 9(3): 315-320
[9] Davies K M, Schwinn K E, Deroles S C, Manson D G, Lewis D H, Bloor S G, Bradley J M(2003)Enhancing anthocyanin production by altering competition for substrate between flavonol synthase and dihydroflavonol 4-reductase. Euphytica131: 259–268
[10] Elomaa P, Honkanen J, Puske R, Sepponen P, Helariutta Y, Mehto M et al (1993) Agrobacterium-mediated transfer of antisense chalcone synthase cDNA to Gerbera hybridainhibits flower pigmentation. Biotechnology 11: 508-511
[11] Fukui Y, Tanaka Y, Kusumi T, Iwashita T and Nomoto K (2003)A rationale for the shift in colour towards blue in transgenic carnation flowers expressing the flavonoid 30,50-hydroxylase gene. Phytochemistry 63: 15-23
[12] Fukusaki E, Kawasaki K, Kajiyama S, An C, Suzuki K, Tanaka Y, Kobayashi A(2004)Flower color modulations of Toreniahybrida by down regulation of chaleone synthase genes with RNA interference. Journal of biotechnology 111:229-240
[13] Gao J Y, Parks C R, Du Y Q(2005)Collected species of the genus camellia an illustrated outline.Hangzhou: Zhejiang Science and Technology Press.90-126
[14] Grotewold E (2006)The genetics and biochemistry of floral pigments. The Annual Review of Plant Biology 57:761-80
[15] Holton TA, BruglieraF, Tanaka Y (1993) Cloning and expression of flavonol synthase from Petunia hybrida. Plant J. 4:1003–1010
[16] Katsumoto Y, Fukuchi-Mizutani M, Fukui Y, Brugliera F, Holton T A,Karan M, Nakamura N et al(2007) Engineering of the rose flavonoid biosynthetic pathway successfully generated blue-hued flowers accumulating delphinidin. Plant Cell Physiol48(11):1589-1600
[17]Lepiniec L, Debeaujon I, Routaboul J M, Baudry A, PourcelL,Nesi N, Caboche M(2006)Genetics and biochemistry of seed flavonoids. Annual Review of Plant Biology 57:405-430
[18]Livak K J, Schmittgen T D(2011). Analysis of relative gene expression data using real-time quantitative PCR and the 2 (-Delta Delta C (T)) method. Methods 25(4): 402-408
[19]Markham KR, Hammett KRW (1994)The basis of yellow colouration in Lathyrusaphaca flowers. Phytochemistry 37:163-165.
[20]Miyajima I, Uemoto S, Sakata Y, Arisumi K, Toki K(1985)Yellow pigment of Camellia chrysantha Flowers. American Camellia year book.71-80
[21]Nielsen K, Deroles S C , Markham K R, Bradley M J, Podivinsky E, Manson D(2002) Antisense flavonol synthase alters copigmentation and flower color in lisianthus. Molecular Breeding, 9:217-229
[22]Nishimoto S, Hashimoto F, Shimizu K, Sakata Y(2004) Petal coloration of interspecific hybrids between Camellia chrysantha and C. japonica. J. Jan.Soc.Hort.Sci.73(2):189-191
[23]Parks C R (2000) Breeding progress with yellow camellia. American Camellia Yearbook, 9-10
[24]Peer W A, Murphy A S (2007) Flavonoids and auxin transport: modulators or regulators? Trends Plant Sci. 12: 556-563
[25]Peng X, Yu D Y, Feng B M, Tang L, Wang Y Q, Shi L Y(2011). Chemical constituents from the Camellia chrysantha. Guihaia 31(4):550-553
[26]Qiu W, Liu M, Qiao G, Jiang J, Xie L, Zhuo R(2011) An IsopentylTransferase Gene Drivenby the stress-inducible rd29A promoter improves salinity stress tolerance in transgenic tobacco. Plant MolBiol Rep. Published online: 23 September 2011
[27]Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B(2007)Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. The plant journal 50:660-667
[28]Stracke R, Jahne O, Keck M, Tohge T, Niehaus K, Fernie A R, Weisshaar B(2010).Analysis of production of flavonol glycosides-dependent flavonol glycoside accumulation in Arabidopsis thaliana plants reveals MYB11-, MYB12- and MYB111-independent flavonol glycoside accumulation. New Phytologist 8(4): 985-1000
[29]Subramanian S, Stacey G, Yu O (2007). Distict, crucial roles of flavonoids during legume nodulation. Trends in plant science 12:282-285
[30]Sun Y, Li H, Wang L S, Dai S L(2011). Rapid, effective method for anthocyanin analysis in tobacco corolla.Chinese Bulletin of Botany46(2):189-196
[31]Tanaka Y, Sasaki N, Ohmiya A(2008).Biosynthesis of plant pigments: anthocyanins, betalains and carotenoids.Plant J54:733-749
[32]Tayler L P, Grotewold E (2005) .Flavonoids as developemental regulators. Current Opinion in Plant Biology,8(3):317-323
[33]To K Y, Wang C K(2006) Molecular breeding of flower color. Floriculture, Ornametal and Plant biotechnology Volume I:300-310
[34]Treutter D(2005). Significance of flavonoids in plant resistance and enhancement of their biosynthesis. Plant Biol. 7:581-591
[35]Tsuda S, Fukui Y, Nakamura N, Katasumoto Y, Yonekura-Sakakibara K, Fukuchi-Mizutani M, et al(2004) Flower color modification of Petunia hybridacommercial varieties by metabolic engineering. Plant Biotechnology, 2004, 21(5): 377-386
[36]van der Krol A R, Lenting P E , Veenstra J, van der Meer I M, Koes R E, Gerats A.G.M. et al(1988) An antisense chalcone synthase gene in transgenic plants inhibits flower pigmentation.Nature 333: 866-869
[37]Winkel-Shirley B (2001) Flavonoid biosynthesis: A colorful model for genetics, biochemistry, cell biology, andbiotechnology. Plant physiology 126: 485-493
Web design by Tribal Systems
